



Spacers are incorporated to introduce flexibility and distance between functional groups and the oligonucleotide hybridization region. These inert molecular linkers reduce steric hindrance, enhancing the efficiency of attached moieties such as dyes or affinity tags.
Key applications include but are not limited to: FRET optimization, structural biology studies, and reduction of steric hindrance.
Spacers & Structural Modifications
Spacers and structural modifications are essential tools for precisely engineering the physical and functional properties of oligonucleotides. These insertions allow you to adjust the distance between functional groups (e.g., a fluorophore and a quencher) or control the structural flexibility of the molecule. This fine-tuning is critical for optimizing interactions, preventing steric hindrance, and achieving desired biological activity or detection efficiency.
Spacers & Structural Modifications | ||
Spacer C3 | Spacer 9 | dSpacer |
Spacer C6 | Spacer 12 | PC Linker |
Spacer C12 | Spacer 18 | |
Spacer C3: A short, 3-carbon chain. Often used as a minimalist spacer or to block the 3'-end of an oligo to prevent enzymatic extension.
Spacer C6: A 6-carbon chain. A very common and versatile length for attaching various molecules (like fluorophores or biotin).
Spacer C12: A longer, 12-carbon chain. Provides more distance for applications where a greater separation is needed.
Spacer 9: Spacer 9 is a triethylene glycol spacer that can be incorporated at the 5'-end or 3'-end of an oligo or internally. Multiple insertions can be used to create long spacer arms.
Spacer 12: A longer PEG-based spacer, providing even more flexibility and distance than Spacer 9/18.
Spacer 18: A 18-atom PEG chain. Excellent for significantly reducing steric hindrance. Ideal for placing between a large dye and the oligo or between biotin and the oligo to ensure efficient binding to streptavidin.
dSpacer: This is a unique spacer that mimics a natural nucleotide missing its base. Primarily used in structural studies to see how enzymes (like those involved in DNA repair) recognize damaged DNA, or to introduce flexibility into a structure.
PC Linker: PC Linker (photocleavable) is a non-nucleosidic molety that can be used to link two nucleotide sequences through a short, UV photo-cleavable C3 spacer arm, this can be added at any position of the sequence.

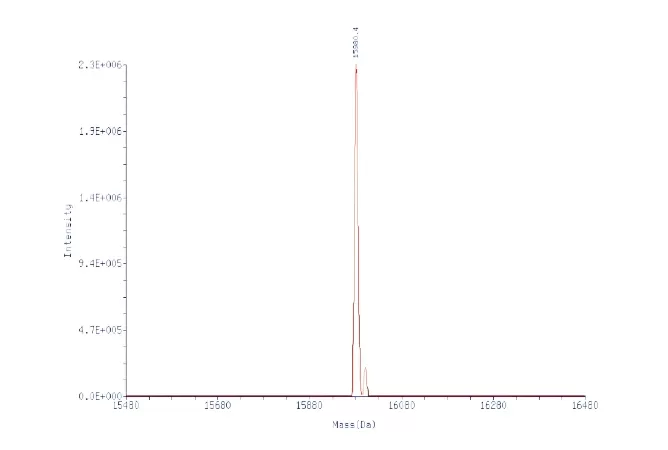
Modification: (dSpacer), 3'Spacer(C3)
Download the order form "Tsingke_DNA_Order Form.1.1.1.250815.csv" for DNA modifications or "Tsingke_RNA_Order Form.1.1.1.250815.csv" for RNA modifications below and email it to info@tsingke.com.cn, or "Send Your Request" to submit your inquiry online. Please refer to "Tsingke_DNA_Modification List_1.1.1.250815.csv" or "Tsingke_RNA_Modification List_1.1.1.250815.csv" sheet to paste special base and internal modification codes in your sequence.
To maintain stability and performance, spacer-modified primers should be stored as follows:
(1) Protect from light: Use opaque or light-blocking containers.
(2) Low temperature: Store at -20°C or below.
(3) Dry conditions: Keep tightly sealed, preferably with a desiccant, to limit moisture and oxidation.
(4) Minimize freeze-thaw cycles: Aliquot upon receipt to avoid repeated freezing and thawing.